Science is sometimes criticized as draining the meaning and beauty out of existence, when in reality it has the opposite problem.
Because it allows one to acquire knowledge otherwise inaccessible to common perception, science gives us access to a whole other landscape, with terrifying and beautiful scenery. It is hard to discuss the history of earth through deep time without falling into epic geopoetry: an oxygen catastrophe so intense the oceans rusted; the sudden diversification of life at the beginning of the Cambrian; the collapse of a habitable Antarctica with the opening of the Drake passage and the start of the circumpolar currents.
I’m going to share a few of the stranger creatures that live in this landscape, or the glimpses that we’ve gotten of them through the haze. They are horrifying, but beautiful, and I think that they make us look at everyday things in new, challenging ways.
“He spoke of lands not far
nor lands they were in his mind.
Of fusion captured high
where reason captured his time.”
Life on land is hard. It’s dry, for one thing; a lot of evolutionary engineering goes into maintaining moisture, and it’s irradiated by ultraviolet light. It’s a really interesting story how complex life spread to the land, but one of the creepier chapters is the evolution of land plants. Did early algae just throw themselves on the beach, slowly evolving in intermediate environments? Some scientists have theorized that something more interesting was happening on those primordial shores.
More than 90% of plants are mycorhizal. That is to say, they have fungal symbiotes in their root systems. These associations are ancient; fossils in the Rhynie cherts show that they were around 400 million years ago. And they’re weird. For one thing, the fungi can siphon off nutrients from the host plant, then feed them to other plants which get their nutrition solely through this exchange. They also probably play a role in gathering mineral nutrition from the soil for their host plant. Some associations are obligate and very specific; this is why orchids can be difficult to grow.
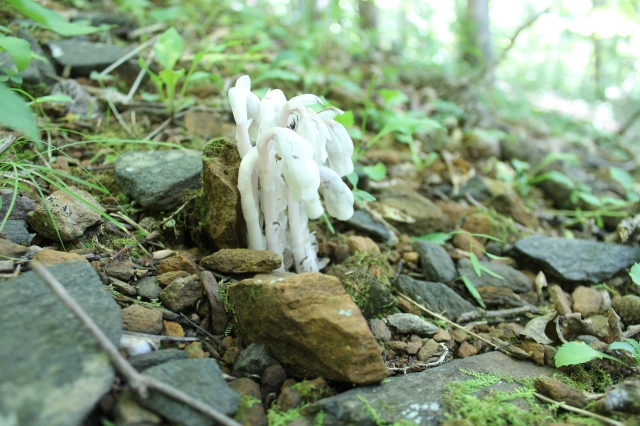

Indian pipes (monotropa) are shown here poking out of the forest floor. They feed exclusively off of a mycorrhizal network which exchanges nutrients with surrounding trees.
What if land plants, what we think of as single organisms, are actually so closely associated with fungi as to be considered partly compose of fungus? That’s the basic idea of the fungal fusion hypothesis. Working together, the fungus and the alga would have been better able to move onto land than either individually. For example, the fungus could have extracted mineral nutrition to the benefit of the alga, while the photosynthetic alga provided food, or compounds to screen the two of them from UV light. Mycorhizal interactions create oily substances in modern plants which protect them from drying out; such an interaction could have protected an early land plant from desiccation as well.
It may have been that land plants were merely an association of fungi and algae, essentially overgrown, vascularized lichens. “Mycotrophism made terrestrial plant life possible,” writes (Pirozynski and Malloch 1975). But it may have been more intimate. Living together, the fungi may well have become an endosymbiont, living inside of the algal cells. Over time, this relationship could have been restructured, fusing fungal genes into a single consolidated genome. Land plants could well be a remix of preexisting creatures. Its not the first such endosymbiosis; chloroplasts and mitochondria arrived on the scene the same way.
Pirozynski KA, & Malloch DW (1975). The origin of land plants: a matter of mycotrophism. Bio Systems, 6 (3), 153-64 PMID: 1120179
Jorgensen R (1993). The origin of land plants: a union of alga and fungus advanced by flavonoids? Bio Systems, 31 (2-3), 193-207 PMID: 8155852
“…It is no lie I can see deeply into the future.
Imagine everything
You’re close
And were you there to stand
So cautiously at first and then so high….“
What is cancer?
Usually, the answer is that it’s cells which have gone on a solo career. They consume the body’s resource and produce nothing useful to the organism, only more of themselves. They are selfishness personified. But there are anomalies which this simplistic account doesn’t explain. For example, why would selfish cancer cells cooperate with one another in tumor formation?
“A longstanding criticism of cancer biology and oncology research is that it has so far taken little account of evolutionary biology,” write (Davies & Lineweaver 2011). They’ve taken a new look through an evolutionary lens and come to the conclusion that cancer is an atavism.
Atavisms are regressions to a more primitive evolutionary states. For example, birds are evolutionarily derived from reptiles with teeth, but do not have teeth themselves. Usually. In rare cases, those ancestral reptilian genes are reactivated, and a bird with teeth will result. That’s an atavism. The notion which these researchers propose is that cancer is an evolutionarily primitive state which emerges when cells are damaged by chemicals, radiation, or time.
The evolution of multicellular life, they propose, required a robust toolkit of cellular communication pathways in order for individuals to cooperate first as a loosely knit colonies with basic division of labor. These tumor-like growths, the first steps toward multicellular life, are what the authors call “Metazoa 1.0“. On top of this basic genetic framework for the self-organization of cells there evolved a further regulatory network responsible for the complex organization of Metazoa 2.0 like us. It’s this delicate regulatory network which goes awry in cancer. At that point, the ancient toolkit is activated and the atavistic mode kicks in.
If true, this framework is a hopeful one. Conventionally, the impressive arsenal of skills available to cancer has been explained as evolutionary adaptations, fed by the cells’ high rate of growth and mutation. This would mean that its arsenal is open-ended, able to evolve dynamically in response to therapy. But if it’s an atavism, though, its toolkit is a finite set limited by what was available to metazoa 1.0.
Davies, P. C. W.,, & Lineweaver, C. H (2011). Cancer tumors as Metazoa 1.0: tapping genes of ancient ancestors Physical Biology, 8 (1) DOI: 10.1088/1478-3975/8/1/015001
“Sound did silence me
leaving no trace.
I beg to leave,
to hear your wonderous stories.”
It’s one thing to find unsettling fossils in the genomes of plants, or in pathologies. It would be something else entirely if there was something strange and beautiful lurking deep inside ourselves.
The human genome is littered with dead viruses. It starts with a retrovirus, something like HIV maybe. It reproduces by inserting its genes into its host’s DNA, where they get expressed by the cell’s transcription/translation machinery. Most of these cells die with the host, but every now and then a germ line cell gets infected, a sperm or egg. This viral hitchhiker then gets passed on to the organism’s progeny. Over time, they’re usually inactivated by mutations, or actively silenced, but they pile up. 8% of the human genome is composed of dead viruses.
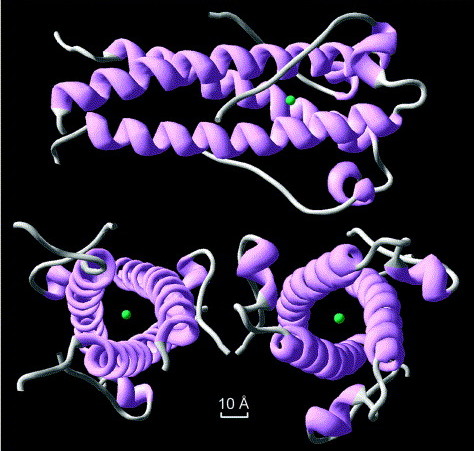
The structure of syncytin, a retrovirus protein which has been recycled in the human genome. Image source from Renard et a 2005; click for link.
It gets weirder. Meet syncytin; it’s a protein which is involved in placental formation in humans. It’s also part of an ancient, repurposed viral gene. Not only was this virus integrated into our genomes, it was rebuilt into an essential part of the mammalian life cycle. It’s happened several times, too: mouse syncytin is derived from a different viral infection than primate syncytin. Experiments on mice show that without this gene, the placenta misforms. Another retrovirus is involved in placenta formation in sheep.
Apparently, mammals across the evolutionary tree have been independently infected by retroviruses, incorporated those viruses into their genomes, and specifically put them to work building placentas. Did these particular viruses just happen to carry genes preadapted for placental formation? Or did mammals evolve and diversify as a result of viral genes flowing into their genomes? What other key biological pathways used to be free-floating viruses?
Mi S, Lee X, Li X, Veldman GM, Finnerty H, Racie L, LaVallie E, Tang XY, Edouard P, Howes S, Keith JC Jr, & McCoy JM (2000). Syncytin is a captive retroviral envelope protein involved in human placental morphogenesis. Nature, 403 (6771), 785-9 PMID: 10693809
Dupressoir, A.,, Vernochet, C.,, Bawa, O.,, Harper, F.,, Pierron, G.,, Opolon, P.,, & Heidmann, T. (2009). Syncytin-A knockout mice demonstrate the critical role in placentation of a fusogenic, endogenous retrovirus-derived, envelope gene. Proceedings of the National Academy of Sciences of the United States of America, 106 (9) DOI: 10.1073/pnas.0902925106


